|
3/20/2024 0 Comments Matlab mutiply vector by 3d matrix Which would be equivalent to the MATLAB m-code: i.e., MTIMESX treats these cases as arrays of 2D matrices and performs the operation on the associated parings. The remaining dimensions are duplicated and specify the number of individual matrix multiplies to perform for the result.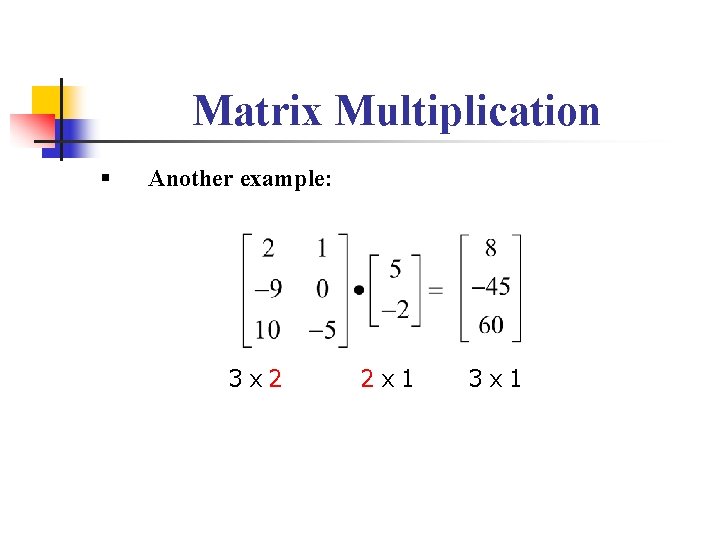 
Mtimesx('SPEEDOMP','OMP_SET_NUM_THREADS(4)') % sets SPEEDOMP mode with number of threads = 4įor nD cases, the first two dimensions specify the matrix multiply involved. 'N' or 'n' = No pre-operation (the default if trans_ is missing)ĭirective = One of the modes listed above, or other directivesĬ = mtimesx(A,B) % performs the calculation C = A * BĬ = mtimesx(A,'T',B) % performs the calculation C = A.' * BĬ = mtimesx(A,B,'g') % performs the calculation C = A * conj(B)Ĭ = mtimesx(A,'c',B,'C') % performs the calculation C = A' * B' Transb = A character indicating a pre-operation on B: 
Transa = A character indicating a pre-operation on A: Where transa, transb, and directive are the optional inputs: The general syntax is (arguments in brackets are optional): SPEEDOMP: Fastest BLAS, LOOPS, or LOOPOMP method even if it doesn't match MATLAB exactly SPEED: Fastest BLAS or LOOPS method even if it doesn't match MATLAB exactly MATLAB: Fastest BLAS or LOOPS method that matches MATLAB exactly (default) LOOPSOMP: Always uses OpenMP multi-threaded C loops if available LOOPS: Always uses C loops if available Can meet or beat MATLAB for speed in most cases Can match MATLAB results exactly or approximately as desired Utilizes BLAS calls, custom C loop code, or OpenMP multi-threaded C loop code Supports Transpose, Conjugate Transpose, and Conjugate pre-operations Supports multi-dimensional (nD, n>2) arrays directly It may be worth having explicit code paths for some lower order matrix dimensions, and in scales well to n-dimensions with the : indexing.MTIMESX is a fast general purpose matrix and scalar multiply routine that has the following features: if you know the dimensions, unrolling the matrix multiply like I did for the two-by-two was an order of magnitude faster. Just FYI here's the Stackoverflow thread where they discussed different approaches and timings. This is presumably faster than the for-loop approach. # Multiply m-by-n matrix with nx1 vector over p slices Multi-dimensional matrix-vector multiplies. However, I think I might be able to rework the function using only It seems like ndmultiply might be robust enough to accomplish this. > to rewrite with bxsfun to be matlab compatible. > but only in Octave since that would use implicit broadcasting. > oh, I forgot it also works if one matrix is 2x2x1 and the other is 2x2xn, > %the multiply, and C will be returned with the intended > %will work for higher dimensions than three. > %otherwise it needs to be the same size as A. > %either A or B will be broadcast in 3rd dimension if it is only a > %will be matrix multiplication of 2x2 matrices along 3rd dimentsion in > %but if both are arrays, they must have the same n. > %either A and B should be 2x2 matrices. It works for any set of 2 x 2 x n x m x p Found a speed comparison somewhere on stackexchange I think. > there is a good vectorized way to do it without knowing the size apriori, > multiply and do vectorized multiplication of each element. 
> fastest way to do it, since my matrix size was fixed, was to 'unroll the > Had to do the 'multiply by slice' problem you mentioned quite a bit. > based program over the summer that worked with n-D arrays of 2x2 matrices. > Are your matrices always a predictable size? I was doing an eigenvector > I am guessing no such routine exists in the core. > is, if A and B are 3-dimensional real arrays, C = multiply_slices(A,B) would > This could be fixed through a _native_ routine to multiply slices. > A triply-nested loop in an octave-financial routine I wrote renders it > On Sun, at 7:21 AM, Nicholas Jankowski On Sun, at 1:24 PM, Nicholas Jankowski wrote: 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
